An integrated bioengineering platform that harnesses the synthetic biology toolbox is the key to an economically viable commercial bioprocess.
Industrial biotechnology enables the replacement of traditional petrochemical processes with benign bioprocesses that harness renewable feedstocks. Such sustainable bioprocesses have the potential to reduce safety risks and overall costs.
Growing concerns over greenhouse gas emissions, volatile fossil energy costs, high capital requirements, and waste from processes using fossil feedstocks (i.e., oil, coal, or natural gas) have inspired scientists and engineers to develop alternative processes that use engineered micro-organisms (1). Microorganisms have the ability to generate products from a variety of renewable feedstocks, such as sugars, biomass, and fats and oils. The feedstock flexibility of microorganisms is a hedge against the volatility of petroleum supplies and prices.
Most of the reaction steps required to convert a feedstock into a desired product (chemical or fuel) can occur inside a microbial host cell, which enables direct production via a single fermentation step. As a result, bioprocesses typically have lower capital costs and risks, smaller environmental footprints, and lower energy consumption than petro-chemical processes.
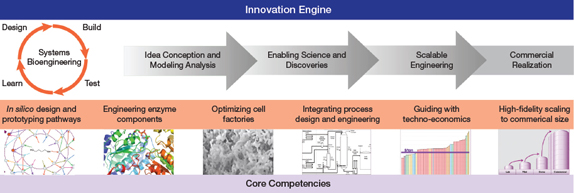
▲Figure 1. This complete platform to industrialize biotechnology includes tightly integrated core competencies across a broad range of disciplines, including computation, experimentation, and bioprocessing.
Genomatica has established an integrated computational and experimental bioengineering platform to design, create, and optimize novel high-producing organisms and bioprocesses (Figure 1) that can compete with traditional petroleum-based processes. The platform covers the entire workflow from idea generation to commercialization. A key feature of the platform is a systems biology approach that enables more-informed and faster decisions, increasing overall efficiency and the speed of strain engineering design-build-test-learn (DBTL) cycles that can be completed.
A broad range of core competencies provides the back-bone of the platform, including in silico design and bio-pathway prediction, optimization of microbial factories using synthetic biology tools, process design, guided techno-economics, and commercial scaling of pathway fermentative processes.
The GENO-BDO™ technology, a sustainable bioprocess for the production of the industrial chemical 1,4-butanediol (BDO) from renewable feedstocks, embodies this platform and whole-process approach best. BDO is a chemical intermediate (with a global market of about two million metric tons) that goes into a variety of products, including plastics manufactured for automobiles, electronics, and apparel. It is currently produced commercially through energy-intensive petrochemical processes using hydrocarbon feedstocks.
A whole-process approach for delivering a commercial bioprocess
Genomatica engineers adopted a process-driven strategy for developing a commercial BDO bioprocess, and envisioned the entire process at full-scale (e.g., feedstock, conversion, separation, etc.) prior to conducting experimental work (2, 3). This approach enabled parallel development and optimization of the engineered microorganism with the overall process.
A key foundation for this approach was a techno-economic analysis (TEA), which generated a complete picture of the total capital investment and production costs. TEA models are generated at the beginning of a project and are used throughout as a guide, enabled by iterative assessment of the project’s technical parameters weighed with economic prioritization.
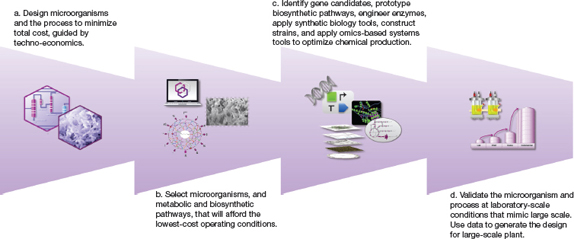
▲Figure 2. A process-driven approach for developing a commercial bioprocess begins with the design of microorganisms, biopathways, and the overall process to minimize operational costs guided by TEA models (a, b). Next, engineers create the production strains and validate their performance under commercially relevant conditions, and generate data for the design of a large-scale plant (c, d).
The first step in this engineering-focused approach is to design the overall chemical production process and microorganism to maximize operation at scale and minimize total costs, guided by the TEA model (Figure 2a). During this stage, multiple organisms are evaluated. Bioengineers use thermodynamics to guide the identification of the best metabolic stoichiometries, fermentation rates, and fermentation conditions, and to select the microorganism, the metabolic pathways, and biosynthetic pathways that will afford the lowest-cost operating conditions (Figure 2b).
The chosen biosynthetic pathways, which encode multiple enzymes that catalyze conversion of a renewable feedstock into the final chemical product, are then engineered into the host microorganism. Bioinformatic analysis is used to identify gene candidates based on information about known enzymes that catalyze similar reactions (4). Introduction of the biosynthetic pathway is facilitated by genetic engineering tools.
Bioengineers typically evaluate the fermentative performance of the engineered cell factory through three essential process metrics: titer (concentration), rate (productivity), and yield (feedstock required to produce a given amount of product), which together are referred to by the acronym TRY. Yield is the most important of the three metrics, as large-scale bioprocesses must operate at 80% or more of theoretical yield to be economically competitive (2). It is not possible for bioprocesses to achieve theoretical yields, because a portion of the carbohydrate feedstock is required for cell growth and maintenance energy. TRY metrics are interdependent and their maximization — through the reduction of byproducts, for example — is key to reducing overall costs.
Next, pathway enzyme engineering and synthetic biology tools are applied iteratively to maximize pathway flux and reduce byproducts (Figure 2c). The systems biology platform uses extensive omics tools — transcriptomics, proteomics, metabolomics, and fluxomics — to probe cellular function.
Transcriptomics and proteomics examine the global expression of genes, while targeted metabolomic approaches analyze pathway intermediates from sugar to product, as well as redox and energy ratios. Fluxomics tools that use 13C-based metabolic flux analysis (5) combine stoichiometric models of metabolism with mass spectrometry methods to trace the conversion of isotopically labeled metabolites. This analysis provides quantitative information on metabolic reactions in vivo and provides the ability to evaluate fluxes through native, engineered, and heterologous pathways.
These tools are applied at laboratory scale to develop and optimize the microorganism and fermentation process under the conditions anticipated at large-scale. The process is then scaled up to generate data for plant design (Figure 2d). The result is a microorganism and fermentation process that translate well to the design of the large-scale production plant with minimal additional capital costs.
Bioengineering a BDO pathway in E. coli
The initial development of the Escherichia coli (E. coli) BDO production strain achieved titers of several g/L BDO from fermentations that lasted nearly five days (6). Because BDO is not a natural chemical, much of the early effort involved the identification of new biochemical pathways to BDO, as well as computational modeling to generate a blueprint of a host strain that would maximize carbon flux to the pathway while balancing overall energy and redox.
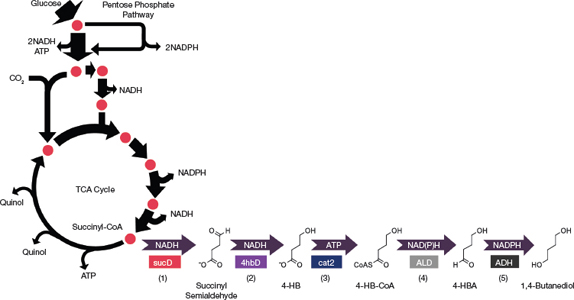
▲Figure 3. A BDO biosynthetic pathway that consisted of five heterologous steps was engineered into E. coli. The enzymes for each numbered step are: 1 = CoA-dependent succinate semialdehyde dehydrogenase (sucD); 2 = 4-hydroxybutyrate dehydrogenase (4hbD); 3 = 4-hydroxybutyrl-CoA transferase (cat2); 4 = CoA-dependent aldehyde dehydrogenase (ALD); 5 = alcohol dehydrogenase (ADH).
Early adoption of the whole-process approach narrowed down the candidate biosynthetic pathways to one that required five heterologous steps to convert the tricarboxylic acid (TCA) cycle intermediate succinyl-CoA into the desired product (Figure 3) (6).
The pathway uses a CoA-dependent succinate semi-aldehyde dehydrogenase (sucD) enzyme to generate succinyl semialdehyde, which is further reduced to 4-hydroxy-butyrate (4-HB) by the 4-hydrobutyrate dehydrogenase (4hbD) enzyme. The 4-hydroxybutyrl-CoA transferase (cat2) enzyme adds CoA to activate 4-HB. The final two reduction steps are catalyzed by the aldehyde dehydrogenase (ALD) and alcohol dehydrogenase (ADH) enzymes to reduce 4-HB-CoA and produce BDO.
The BDO pathway is...
Would you like to access the complete CEP Article?
No problem. You just have to complete the following steps.
You have completed 0 of 2 steps.
-
Log in
You must be logged in to view this content. Log in now.
-
AIChE Membership
You must be an AIChE member to view this article. Join now.
Copyright Permissions
Would you like to reuse content from CEP Magazine? It’s easy to request permission to reuse content. Simply click here to connect instantly to licensing services, where you can choose from a list of options regarding how you would like to reuse the desired content and complete the transaction.
